Congratulations to Pia!
Pia just received her MSc in Cell Biology from ETH Zurich in November with her MSc thesis "CRISPR-KO screening for regulators of PAX5 abundance in B-ALL".
Pia just received her MSc in Cell Biology from ETH Zurich in November with her MSc thesis "CRISPR-KO screening for regulators of PAX5 abundance in B-ALL".
Magda is a Ph.D. student at the Institute of Bioorganic Chemistry Polish Academy of Sciences. In this academic year, she received the Etiuda 8 grant which is...
Magda is a Ph.D. student at the Institute of Bioorganic Chemistry Polish Academy of Sciences. In this academic year, she received the Etiuda 8 grant which is dedicated to young scientists who want to do an internship abroad. Magda has been using CRISPR-Cas9 technology to shorten mutated CAG repeat tract in genes associated with polyQ disorders. Her scholarship is scheduled for 6 months.
Congratulations to our Postdoc Research Fellow, Matthias Muhar, who received the Marie Sklodowska-Curie Actions (MSCA) individiual fellowship which belongs to the EU Horizon 2020 program.
Eric joined the Corn lab as a Postdoc in November 2020. He has received his PhD in Biochemistry, Molecular Biology, and Biophysics from the University of Minnesota...
Eric joined the Corn lab as a Postdoc in November 2020. He has received his PhD in Biochemistry, Molecular Biology, and Biophysics from the University of Minnesota in 2020 in the lab of Dr. Wendy Gordon. He is interested in studying synthetic lethalities and viabilities in DNA repair utilizing CRISPR.
Roman joined the Corn lab in October 2020. Earlier this year he has received his Ph.D. from the University of Boulder, where he had developed tools by combining...
Roman joined the Corn lab in October 2020. Earlier this year he has received his Ph.D. from the University of Boulder, where he had developed tools by combining CRISPR and aptamer components in the group of Prof. Batey. He is currently interested in finding new ways to understand DNA damage repair and developing new tools to easily modify DNA.
Susanne joined the Corn Lab in September 2020 to support the team with High Throughput sequencing applications. She received her PhD in Paleogenomics at the...
Susanne joined the Corn Lab in September 2020 to support the team with High Throughput sequencing applications. She received her PhD in Paleogenomics at the University of Mainz. Susanne has a long history in Next Generation Sequencing, starting with sequencing of ancient genomes. After working in a clinical setting, she supported several research groups of the University/ETH Zurich in their genomics and transcriptomics applications using Illumina workflows. Susanne will be working at GEML implementing new protocols and sequencing strategies in genome editing workflows.
John received his PhD in Oncology from the University of Oxford in 2020, working with Prof. Kristijan Ramadan on the molecular mechanisms of DNA-protein crosslink...
John received his PhD in Oncology from the University of Oxford in 2020, working with Prof. Kristijan Ramadan on the molecular mechanisms of DNA-protein crosslink repair. He joined the Corn Lab as a postdoctoral researcher in October 2020. His research interests include multiplexed genome engineering, DNA repair, and synthetic lethality.
When using CRISPR genome editing in stem cells, it’s far easier to break a gene with indels than to fix it with HDR. This manifests in an interesting way. If you...
When using CRISPR genome editing in stem cells, it’s far easier to break a gene with indels than to fix it with HDR. This manifests in an interesting way. If you monitor a “CD34+” population of hematopoietic stem and progenitor cells (HSPCs) from the bone marrow, indels start high and stay high but HDR alleles are lost over time. Why do these different genetic outcomes differ over time? Is HDR bad for the long-term stem cells? Or is editing in the CD34+ population actually heterogeneous, and different cells get different alleles? New work from postdoc Jenny Shin in the lab, out in Cell reports, both answers this question and finds a way to fix the problem.
Jenny and collaborators used a powerful combination of immunophenotyping, next generation sequencing, and single-cell RNA-sequencing to investigate and reprogram genome editing outcomes in subpopulations of adult human CD34+ HSPCs. These HSPCs are actually several different types of cells, including more differentiated progenitors that cycle and very “stemmy” long-term HSCs that are quiescent. The team found that there is a dramatic tension between HDR and quiescence in LT-HSCs. Quiescent stem-enriched cells utilize NHEJ and exhibit almost no HDR. By contrast, non-quiescent cells with the same immunophenotype utilize both NHEJ and HDR. Quiescence is critical for engraftment and stem cell maintenance, so it was now clear that all cells in the CD34+ population get indels and the cycling progenitors were getting HDR alleles, but the quiescent LT-HSCs weren’t doing HDR.
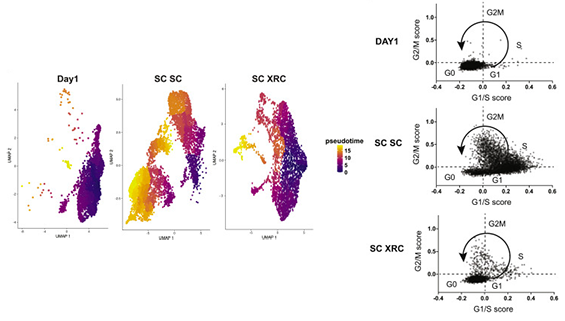 Jenny then had a very creative idea. She asked if a previously reported small molecule cocktail, “XRC”, that maintains quiescence could be used after the fact to re-quiesce LT-HSCs. Using this new strategy and good timing, she found a way to get LT-HSCs with high levels of HDR by briefly allowing them to cycle during editing, and then inducing quiescence later on. This yielded a 6-fold increase in the HDR/NHEJ ratio in quiescent stem cells ex vivo and during long-term engraftment in mouse experiments. The re-quiescence strategy might in future be combined with engineered Cas9-geminin constructs that reduce NHEJ, further tipping the balance towards HDR. Jenny’s results highlight the tradeoffs between editing and fundamental cellular physiology and suggests strategies to manipulate quiescent cells for research and therapeutic genome editing.
Jenny then had a very creative idea. She asked if a previously reported small molecule cocktail, “XRC”, that maintains quiescence could be used after the fact to re-quiesce LT-HSCs. Using this new strategy and good timing, she found a way to get LT-HSCs with high levels of HDR by briefly allowing them to cycle during editing, and then inducing quiescence later on. This yielded a 6-fold increase in the HDR/NHEJ ratio in quiescent stem cells ex vivo and during long-term engraftment in mouse experiments. The re-quiescence strategy might in future be combined with engineered Cas9-geminin constructs that reduce NHEJ, further tipping the balance towards HDR. Jenny’s results highlight the tradeoffs between editing and fundamental cellular physiology and suggests strategies to manipulate quiescent cells for research and therapeutic genome editing.
Pia received her Bachelor’s degree in Biology from ETH Zürich in 2018. She joined the Corn Lab as a Master student in February 2020 working on a CRISPR-based ...
Pia received her Bachelor’s degree in Biology from ETH Zürich in 2018. She joined the Corn Lab as a Master student in February 2020 working on a CRISPR-based screen for regulators in B-cell leukemia. Her research interests include genome engineering, functional genomics, cancer, and DNA repair mechanisms.
Questions and/or comments about Corn Lab and its activities may be addressed to: