Pushing Cas9 off the genome, published in Molecular Cell
Cas9 is a great DNA cutting enzyme, but it’s also a little weird. Unlike other nucleases (such as restriction enzymes), S, pyogenes Cas9 sticks on DNA for a looooong time. In fact, it spend the same amount of time on DNA whether it is active or inactive! In a test tube, it takes hours and hours to let go of even a cleaved DNA strand. So how does genome editing even work? Does a human cell care that Cas9 it stuck on its genome? Does it have ways to knock it off?
Superstar grad student Alan Wang’s new paper, out today in Molecular Cell, solves this mystery. Previous work suggested that RNA polymerase II was capable of displacing Cas9 in vitro. But Cas9-based technologies work even when targeted to a non-transcribed region. Alan’s first hint that something interesting was going on came from work in Xenopus egg extract, in collaboration with the lab of Johannes Walter. This “frog juice” is very often used to study DNA repair and can be elegantly deconstructed to figure out biological function. Cas9 in buffer sticks on DNA for a long time, but in Xenopus extract it comes off almost immediately!
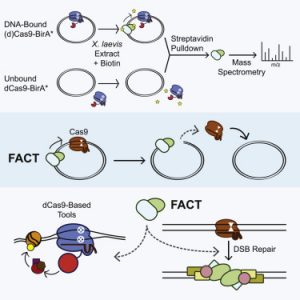 Alan took an unbiased approach to figure out what was removing Cas9 from DNA. He fused recombinant Cas9 with a promiscuous biotin ligase, bound purified Cas9-ligase to a plasmid, and used mass spectrometry to figure out what pushed the Cas9 off the plasmid. We were pleasantly surprised to find that both subunits of a dimeric histone chaperone called FACT were top mass spec hits! Follow-up experiments showed that FACT was necessary and sufficient for displacing Cas9 from DNA substrates. FACT was responsible for turning Cas9 from a multi-turnover “classic” nuclease enzyme into a single-turnover sticky enzyme!
Alan took an unbiased approach to figure out what was removing Cas9 from DNA. He fused recombinant Cas9 with a promiscuous biotin ligase, bound purified Cas9-ligase to a plasmid, and used mass spectrometry to figure out what pushed the Cas9 off the plasmid. We were pleasantly surprised to find that both subunits of a dimeric histone chaperone called FACT were top mass spec hits! Follow-up experiments showed that FACT was necessary and sufficient for displacing Cas9 from DNA substrates. FACT was responsible for turning Cas9 from a multi-turnover “classic” nuclease enzyme into a single-turnover sticky enzyme!
In human cells, FACT had several interesting effects on genome manipulation. Knocking down FACT delayed homology directed repair and altered the balance of repair outcomes. FACT knockdown increased epigenetic marking from both CRISPRi and CRISPRa constructs, and increased CRISPRi phenotypes. We attribute this to increasing the residence time at a target site: giving the effect fused to Cas9 more time to have an effect on the genome.
The take-home is that cells are not passive players, and play a leading role in genome manipulation. The cell is just as important as the enzyme! Alan’s work starts to reveal how cells monitor their genomes during Cas9 interventions.


